CRISPR Gene Editing
CRISPR Gene Editing
Related Tools
Related Labs
Related Worksheets
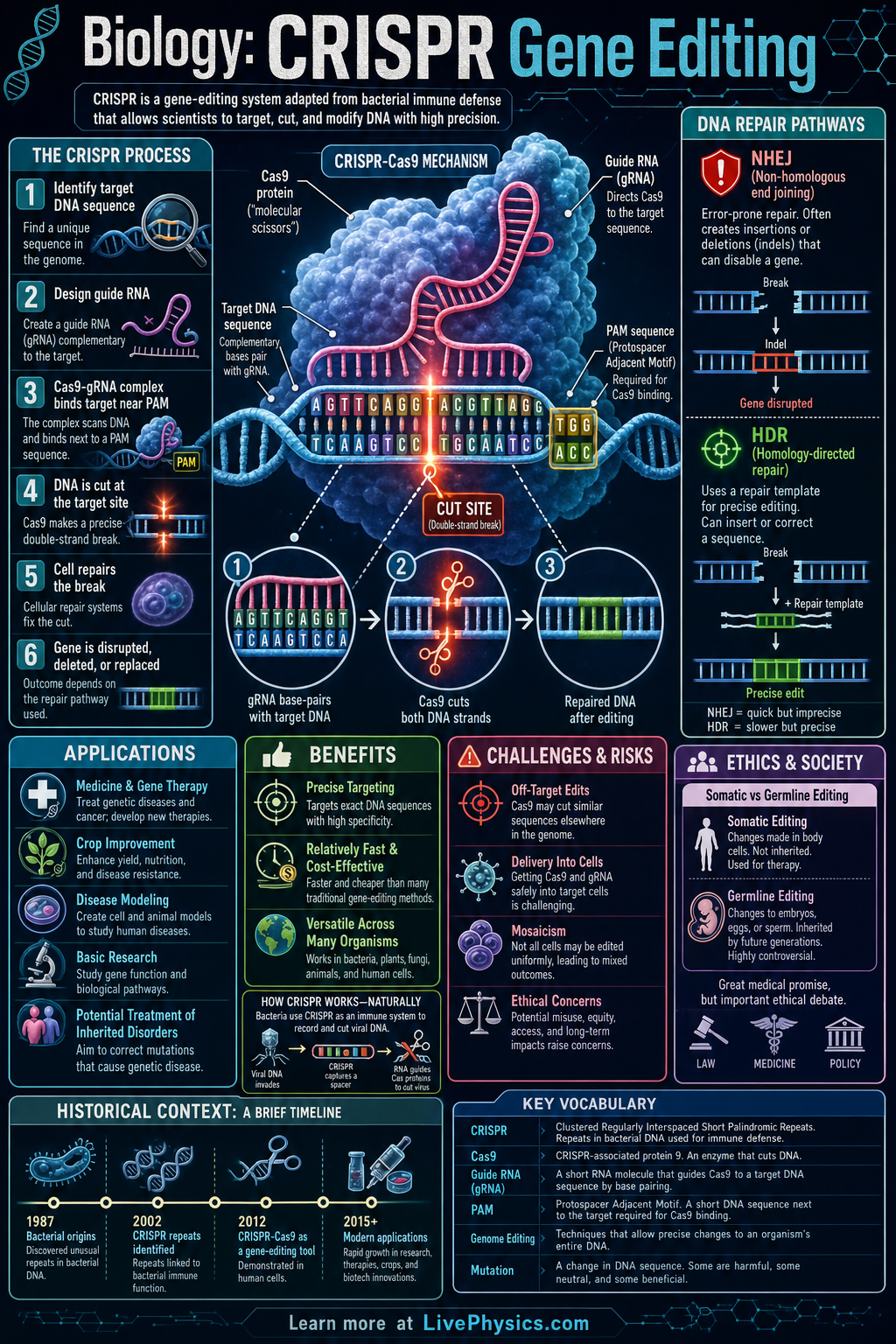
CRISPR gene editing is a powerful biotechnology that lets scientists change DNA at chosen locations. It matters because DNA controls how cells function, so editing a gene can help researchers study disease, improve crops, and potentially treat inherited disorders. The most widely known version uses the Cas9 protein together with a guide RNA to find a matching DNA sequence. This has made gene editing faster, cheaper, and more precise than many older methods.
CRISPR works by using a guide RNA that pairs with a target DNA sequence next to a short PAM sequence. The Cas9 enzyme binds there and cuts both strands of DNA, creating a double strand break. A cell then repairs the break either by nonhomologous end joining, which often disrupts the gene, or by homology directed repair, which can insert or replace a chosen DNA sequence. The outcome depends on the target, the repair pathway, and how accurately the guide RNA matches the DNA.
Key Facts
- CRISPR stands for Clustered Regularly Interspaced Short Palindromic Repeats.
- Cas9 uses a guide RNA to recognize a complementary DNA target sequence.
- A target site usually must be next to a PAM sequence such as NGG for SpCas9.
- After Cas9 cuts DNA, cells repair the break by NHEJ or HDR.
- NHEJ often creates small insertions or deletions called indels that can knock out a gene.
- Gene editing accuracy depends on guide RNA matching, PAM presence, and limiting off target cuts.
Vocabulary
- Guide RNA
- A short RNA sequence that directs Cas9 to a matching DNA target.
- Cas9
- An enzyme that cuts DNA at the location specified by the guide RNA.
- PAM
- A short DNA sequence next to the target that Cas9 must recognize before cutting.
- NHEJ
- Nonhomologous end joining is a repair process that reconnects broken DNA and often introduces small errors.
- HDR
- Homology directed repair is a repair process that uses a template to make a precise DNA change.
Common Mistakes to Avoid
- Thinking CRISPR always makes a perfect edit, which is wrong because cells repair cuts in different ways and off target changes can occur.
- Ignoring the PAM requirement, which is wrong because Cas9 cannot cut a target unless the correct PAM is nearby.
- Assuming the guide RNA binds any similar sequence, which is wrong because mismatches can reduce cutting or cause unwanted off target binding.
- Confusing NHEJ with HDR, which is wrong because NHEJ usually creates disruptive indels while HDR can copy in a designed sequence.
Practice Questions
- 1 A DNA target region reads 5'-TACGGAACCTGG-3' and the required PAM for SpCas9 is NGG. Identify one possible PAM in this sequence and state whether Cas9 could potentially target a nearby site.
- 2 In an experiment, 200 cells receive CRISPR-Cas9. If 55% are edited by NHEJ and 15% are edited by HDR, how many cells show NHEJ edits and how many show HDR edits?
- 3 A scientist wants to disable a gene completely rather than insert a new DNA sequence. Explain why NHEJ is often more useful than HDR for this goal.